Tatyana Shelkovnikova Lab
RNA metabolism in health and disease: mechanisms and drug discovery

RESEARCH
We have several fundamental and translational projects falling into three main themes
1) Molecular mechanisms of amyotrophic lateral sclerosis (ALS) and related neurodegenerative disorders 2) Regulation of stress granules and paraspeckles under stress and in disease states 3) Therapeutic targeting of RNA
Research Projects
Amyotrophic lateral sclerosis (ALS), the most common form of motor neuron disease, is a severe and inevitably fatal neuromuscular condition which currently has no cure. Disease-modifying therapies are urgently needed. To be able to find such therapies, we need better understanding of disease mechanisms, especially those applicable across different disease subtypes. Our research is focusing on the components of cellular RNA metabolism dysregulated in ALS, including lncRNA NEAT1 and RNA-binding proteins such as FUS and TDP-43.
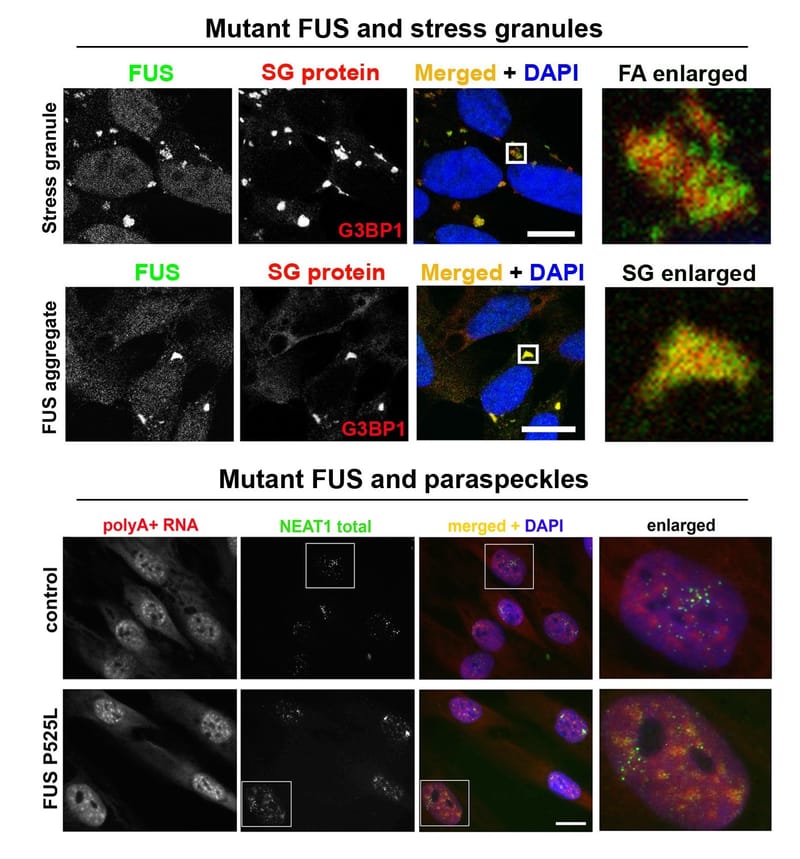
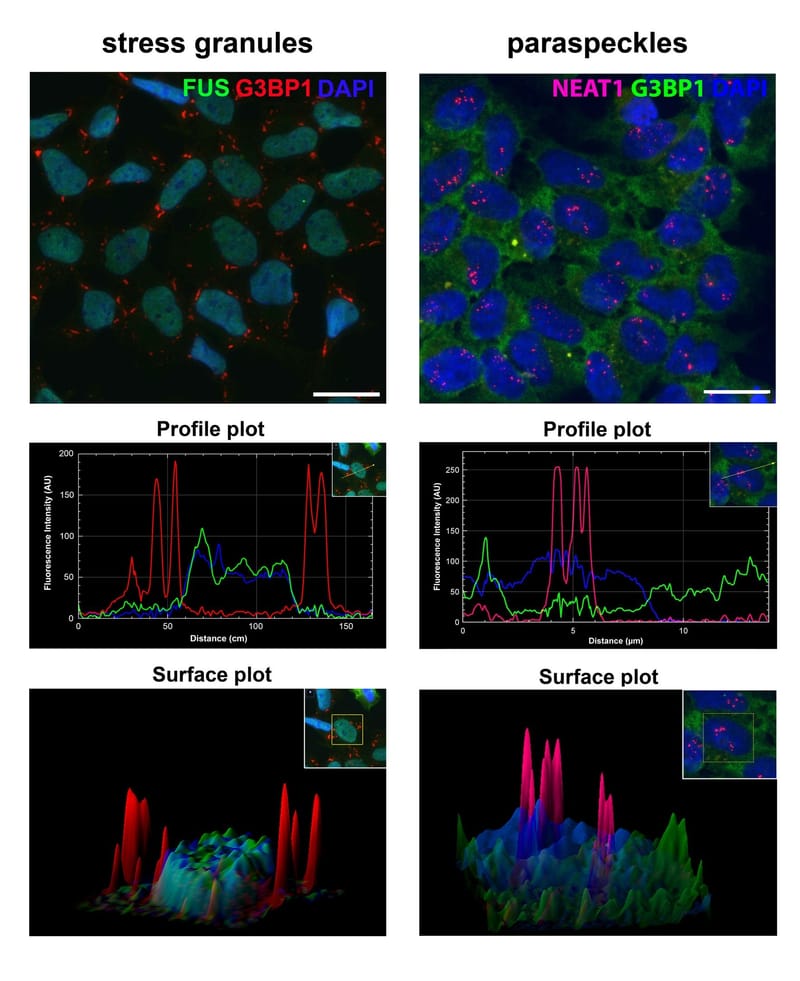
Read MoreStress granules and paraspeckles are prototypical RNP granules localised in the nucleus and cytoplasm, respectively. Recently, we found that despite their spatial separation, these two RNP granules are intimately linked. Our aim is to dissect molecular mechanisms behind paraspeckle-stress granule crosstalk.
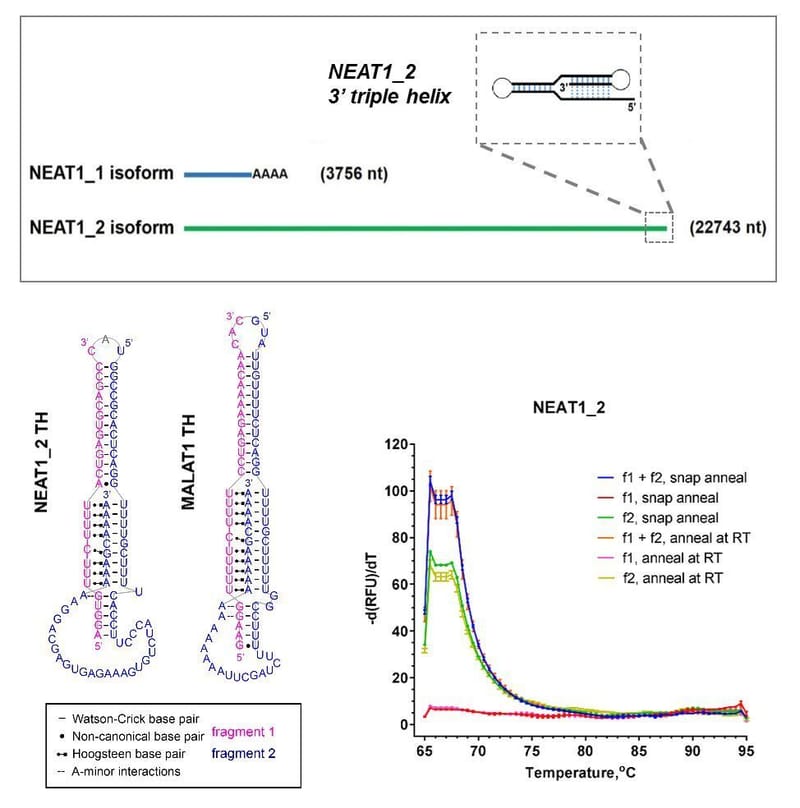
Read MoreMany RNA molecules represent attractive drug targets. However, RNAs are traditionally thought to be difficult to target using small molecules. It is now increasingly appreciated that despite linearity and limited structural complexity, most RNAs contain complex 3D motifs in their structure that can serve as platforms for small molecule binding. We are exploring the possibility of using small molecules for targeting of protein-coding and non-coding RNAs to expand the boundaries of the druggable genome.
Read MoreWho are we?
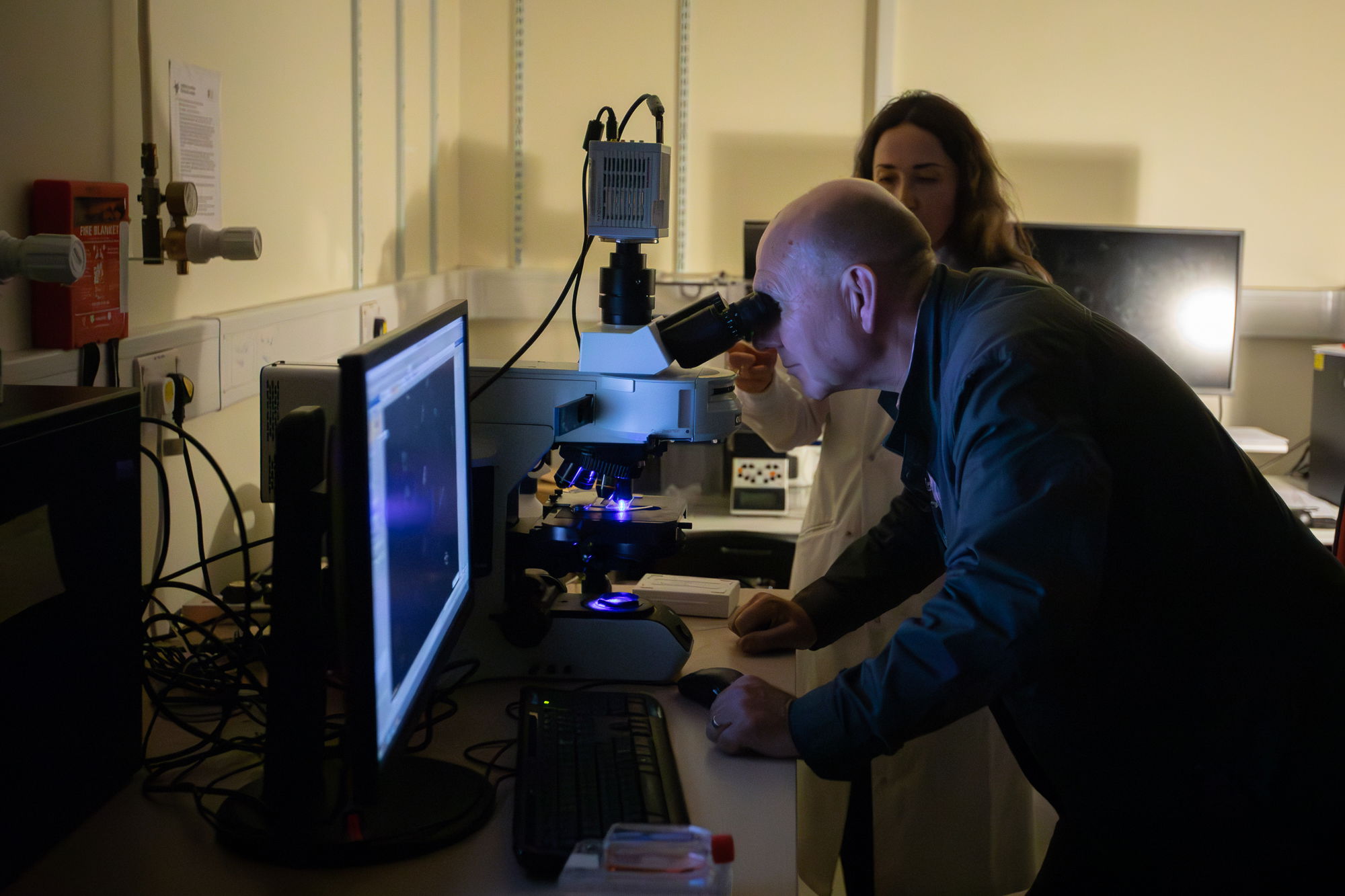
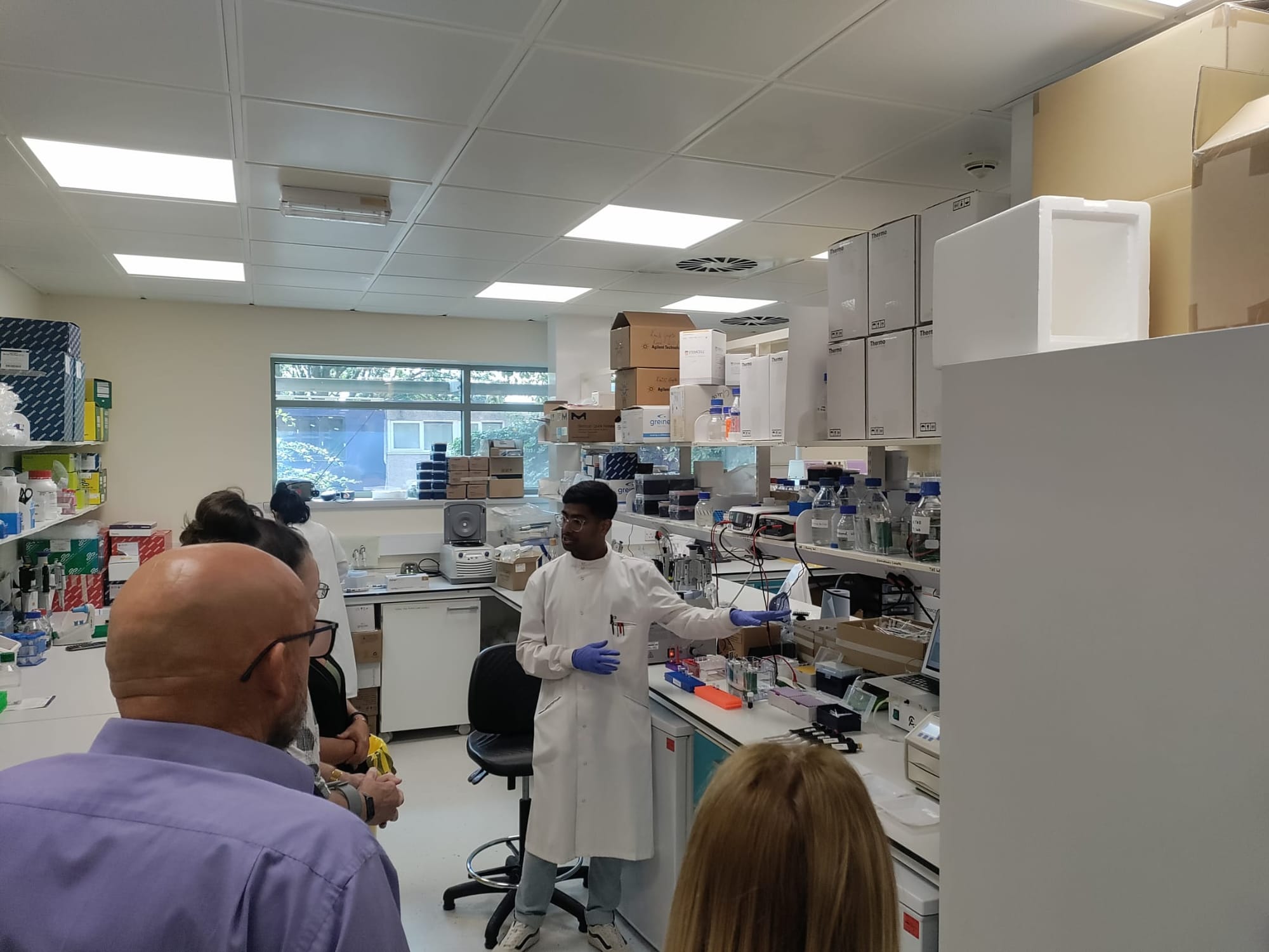
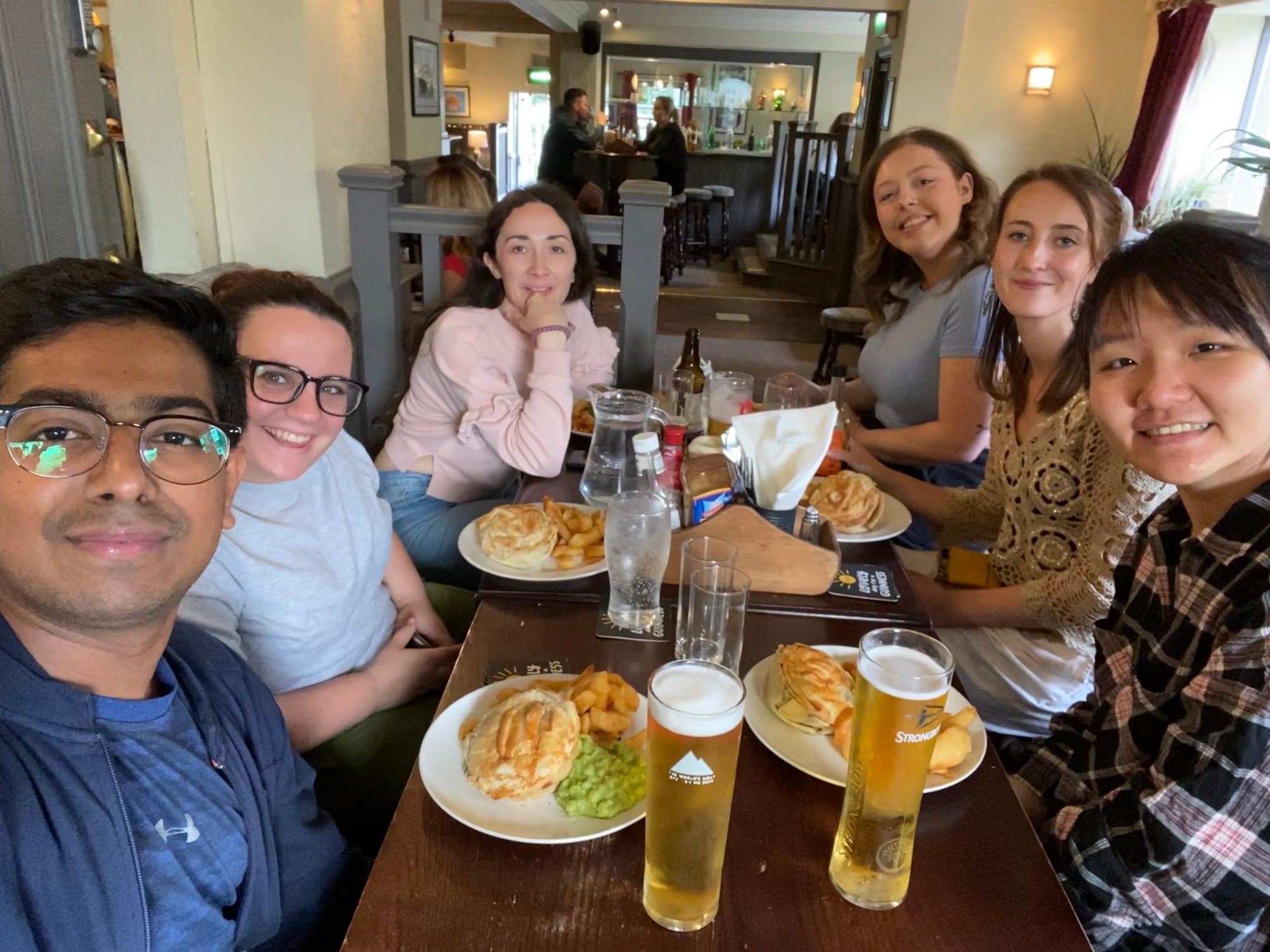
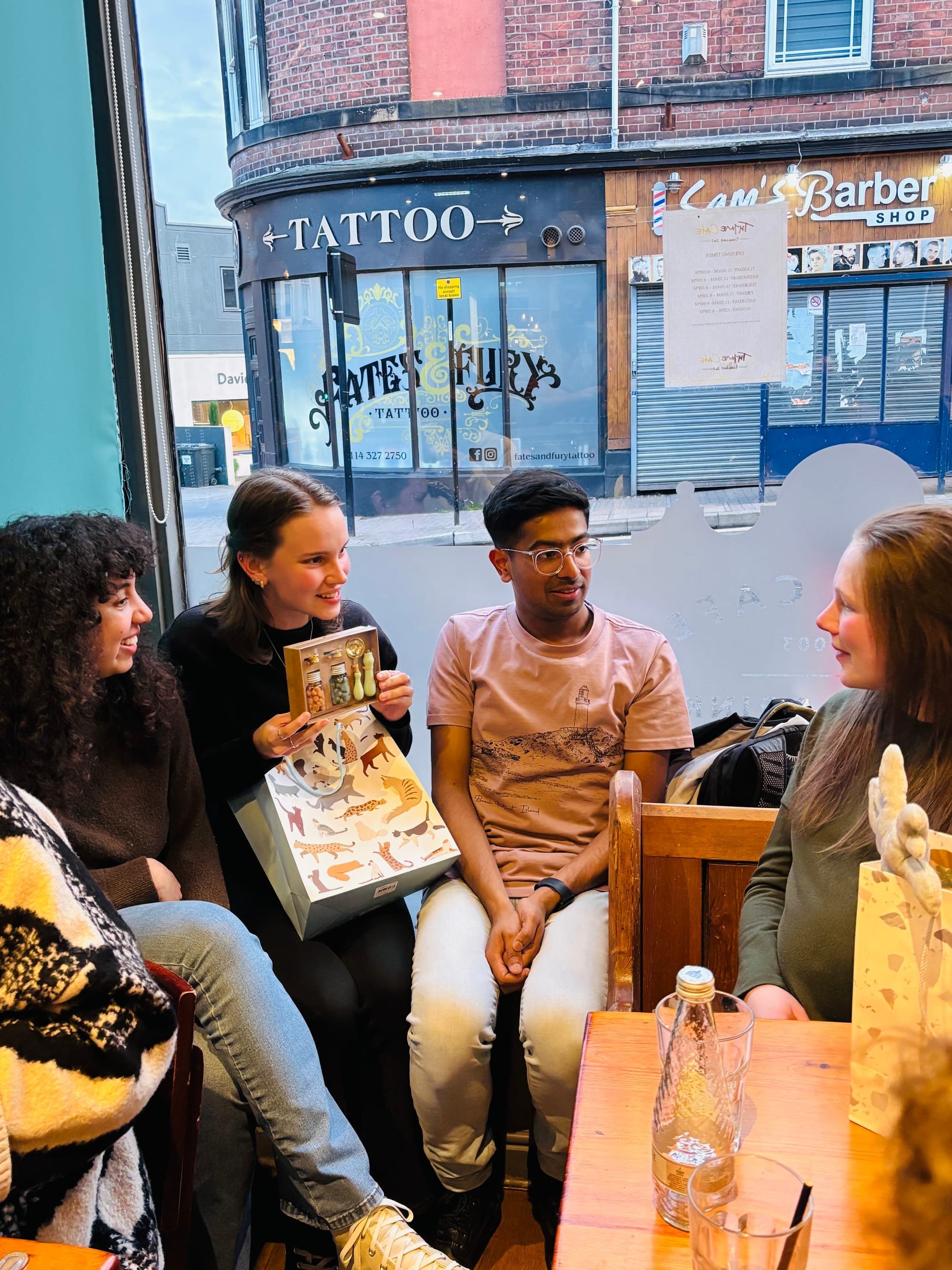
Meet the team
Team
Dr Tatyana Shelkovnikova
Group LeaderI have been involved in biomedical research since 2008. My PhD project was aimed at development and validation of in vitro and in vivo models for testing the effects of neuroprotective compounds. In 2012, after completion of PhD studies and a short-term EMBO fellowship, I joined Prof Buchman’s laboratory at Cardiff University as a postdoctoral researcher. Between 2012 and 2015, I worked on the molecular mechanisms of Parkinson’s disease and amyotrophic lateral sclerosis (ALS). Wishing to focus on the previously overlooked link between a nuclear RNA granule, the paraspeckle, and ALS pathogenesis, in 2015 I obtained dedicated funding from the Medical Research Foundation (3-year fellowship). In September 2018, I started as a MND Association Senior Non-Clinical Fellow and independent group leader at the Cardiff University Medicines Discovery Institute. In September 2021, I moved to the Sheffield Institute for Translational Neuroscience (SITraN) and in August 2022, started as a UKRI Future Leaders Fellow. The lab has been fully operational since Nov 2022, when the first lab members were recruited. The lab is currently supported by URKI, MRC, MND Association, BBSRC, MND Scotland and ARUK.
Brittany Ellis
Research AssistantMotivated by my fascination with clinical research from a young age, I chose to study my bachelor’s degree in Biomedical Science. During my degree I completed an NHS placement year in the Immunology lab of the Pathology department at James Cook University Hospital. My final year project focussed the role of folate depletion during pregnancy and gestation on infants’ epigenetic profile of the translocator protein gene. I completed my master’s degree in Stem Cell and Regenerative Medicine at the University of Sheffield investigating Synaptotagmin protein isoforms’ role in modulating exocytosis by G-protein coupled receptors. My most recent year of research has been in the Tsakiridis Lab in the Centre for Stem Cell Biology working on the international CONNECT project (differentiation of enteric nervous system components for a CHIP modelling system aiming to model the Parkinson’s patient nervous system for the assessment of novel therapeutics). My PhD project with the TS group was focused on characterising STMN2 regulation under stress and establishing tools for STMN2-focussed drug discovery. I have now started as a RA on a project funded by My name'5 Doddie, aimed at developing NEAT1/paraspeckle targeted therapies for ALS.
Rachel Hodgson
Postdoctoral researcherI completed an undergraduate degree in Biomedical Science, followed by an MSc in Molecular and Cell Biology. After which, I was keen to pursue a career in academic research and joined the Campbell/Allen lab group at Sheffield Hallam university. My PhD research worked towards characterising the cellular localisation of the protein eIF2B in glial cells, with a focus on regulation by cellular stress response pathways. I then went on to complete a postdoctoral research associate position, investigating how mutations in eIF2B that are causative of the childhood neurodegenerative disease, leukoencephalopathy with vanishing white matter, impact upon the function and cellular localisation of eIF2B. I have recently joined the TS lab at SITraN as a postdoctoral research associate.
Wan-Ping (Cara) Huang
Postdoctoral researcherIn 2019, I gained my PhD degree at the National Chung-Hsing University in Taiwan. My PhD research project focused on studying ribosomal reading-frame switch mechanisms, which is regulated by multiple translational factors such as mRNA secondary structures, charged-tRNAs availability and cellular protein factors. After graduation, I worked in the lab as a postdoctoral researcher for an international collaboration project with Dr. Guillaume Hautbergue to investigate the pathological translation mechanism in C9ORF72-ALS/FTD. It brings me to extend my RNA and translation knowledge and lab skills in the research of neurovegetative diseases. In 2021, I worked in Dr. Guillaume Hautbergue’s group as a postdoctoral researcher in ARUK project to develop compound screening assays for C9ORF72-ALS/FTD. From November 2022, I joined Dr. Tatyana Shelkovnikova’s lab for mechanistic studies of ALS-causative mutations and drugs discovery through an in vitro reconstituted-RNP complexes in MRC grant.
Anna Avila Sanchez
Postdoctoral researcherI studied biomedical sciences in Barcelona, where I’m from. I then moved to Scotland to pursue a master's in integrative neuroscience at Edinburgh University, where I did my project with Dr Matt Livesey performing patch clamp in iPSC-derived neurons to study neurotransmitter-gated ion channel dysregulation in ALS. I have now finished my PhD under Dr Chris Henstridge’s supervision at the University of Dundee, which centres around assessing ALS neuropathology and its association with cognitive impairment. I have experience using high-resolution imaging techniques as well as an ample histological and molecular biology. In the TS lab, I am focusing on biomolecular condensate and RBP dysfunction in MND.
Ruaridh Lang
PhD studentMy name is Ruaridh I have recently graduated from MBiolSci Molecular Biology in the School of Biosciences here at Sheffield. I have since gained a keen interest in understanding disease pathology and subsequent development of therapeutics. Outside of education I enjoy my sports where I have played rugby and competed in athletics for many years. I also enjoy hiking and climbing when the weather is behaving itself! I look forward to meeting everyone and getting started with some science!
Vedanth Kumar
PhD studentHi I am Ved and I was a technician and now PhD student in TS lab. I’ve been part of the lab since April 2022 (as a student in MSc Translational Neuroscience and then as a technician). One of my projects was looking at a novel autoregulatory mechanism of FUS which is mutated protein in subset of Amyotrophic Lateral Sclerosis (ALS) cases. Another project focused on lncRNA semi-extractability and paraspeckles. It has been a very rewarding experience working on these projects and I am delighted to come back to TS lab to do a PhD, funded by the Neuroscience Institute.
Georgia Boothe
PhD studentI studied my undergraduate MSci Neuroscience degree at the University of Southampton. During my studies at university, I took a strong interest in understanding the structural and molecular underpinnings of neurodegenerative diseases and undertook many lab projects investigating protein dysfunction in Alzheimer’s and Parkinson’s disease. My PhD project when joining the TS lab will revolve around the optogenetic modelling of a hallmark neurodegeneration pathology for mechanistic research and drug discovery.
Emily Day
PhD studentI completed my undergraduate degree in Human Physiology at the University of Leeds, where my final year project studied changes in neuronal morphology within the hippocampus of cognitively enhanced PDE4BY358C mice. Following this, I worked as a Research Assistant for 2 years at the University of Oxford, studying genetic regulation of growth in Physcomitrium patens and conducting behavioural assays to study aggressive behaviours in Drosophila melanogaster. I then completed my MSc (Translational Neuroscience) at SITraN, where I undertook my research project within the TS lab, analysing aggregate composition and m6A RNA modification in FUS-linked ALS. I am interested in RNA and protein dysregulation underlying neurodegenerative diseases, with my PhD project looking at the role of post-translational modifications in regulating biomolecular condensates.
Sabin John
Research technicianHi, I’m Sabin! I graduated from the Integrated Master of Pharmacology course at the University of Bath in 2024. My interest in neuroscience began during my year-long placement at Virginia Commonwealth University, where I utilised molecular biology and electrophysiology to study the pathology of Alzheimer's Disease. Now, I am excited to be joining the TS lab as a Research Technician, where I will be working on modelling C9orf72-derived dipeptide-repeat protein pathology for mechanistic ALS/FTD studies!
Katie Gelder
Postdoctoral researcherHi, my name is Katie and I am currently a postdoc in the Shelkovnikova Lab. I was awarded my PhD at the University of Sheffield in 2024, for work done in the Bose Lab in the School of Biosciences. My PhD work focused on studying phase separation, and how intrinsic disorder regions regulate the condensate formation of transcriptional coactivator protein CREB binding protein (CBP). I furthered my work in the Bose Lab as a postdoc where I started to develop single molecule assays to understand how CBP’s activity is changed in different phase separated states. I enjoy learning about how condensates affect protein and RNA function, and how this is related to disease. This is why I am really excited to be working in the Shelkovnikova Lab, where my project will be focused on trying to use techniques such as Cryo-Electron Tomography (Cryo-ET) and Cryo-Correlative Light and Electron Microscopy (Cryo-CLEM) to look at the structure of paraspeckles.
Abbie Martin
PhD studentI completed my undergraduate degree in Biological and Biomedical Sciences with a moderatorship in Microbiology from Trinity College Dublin in 2022, where I studied the underlying molecular mechanisms of Acinetobacter baumannii pathogenesis. Following this, I spent a number of years working in R&D, quality control and regulatory roles in a variety of areas, including infection prevention, water purification, beverage production and high throughput diagnostics. I moved to Sheffield in 2024 to undertake an MSc in Translational Neuroscience, where I completed a six-month project investigating a novel proteinopathy in C9ORF72 ALS under supervision of Prof Guillaume Hautbergue and Dr Robin Highley. I have joined the TS lab as a PhD student, and my project focuses on deciphering the aggregation of dipeptide repeat proteins associated with C9ORF72 pathology via optogenetic and in vitro analyses of modifiers.
Alumni
Previous lab members: Dr Haiyan An (postdoc, 2019-2021), currently scientific officer, Swansea University Medical school. Camille Rabesahala de Meritens (technician, 2019-2021), currently PhD student, University of Geneva. Jana Kelsall (MRes student, 2020-2021). Gioana Litscher (summer student 2019), currently PhD student, University of Zurich. Jacqui Chalakova (research assistant, 2022-23), currently PDRA, King's College London. Emily Day (MSc student, 2023). Jessica Rayment (MSc student and then technician, 2022-23).
Our Supporters
Look into those who make it possible
Funding

- My Name'5 Doddie Foundation
- Alzheimer's Research UK (ARUK)
- UKRI
- Motor Neurone Disease Association
PhD studentship - student to be recruited in 2026
- MRC
- MND Scotland
- BBSRC
- MRC/AZ Centre for Lead Discovery award for the identification of small molecules modulators of NEAT1_2/paraspeckles (co-I, with Cardiff)
Past funding:
- MND Association Senior Non-Clinical Fellowship “Long non-coding RNA NEAT1 in motor neurone physiology and pathophysiology” (2018-2024)
- Medical Research Foundation fellowship “Dissecting the role of the paraspeckle in amyotrophic lateral sclerosis” (2015-2018)
- Academy of Medical Sciences Springboard Award: “Developing tools to monitor expression of lncRNA NEAT1 isoforms for basic research and drug discovery” (2019-2021)
- ISSF Wellcome Trust Translational Kickstart Award “Identification of chemotypes selectively binding to a structured motif of lncRNA NEAT1 as templates for novel drugs” (2019-2021)
- Welsh Government Sêr Cymru III – Tackling COVID-19 grant (2020-2021)
Publications
News
TS presented the NCB paper work at the RNA society meeting 2026 in Montreal, Canada (RNA condensates session). It was fantastic to attend the keynote by Archa Fox reflecting on 25 years of paraspeckle research, including featuring some of our work!
30 March 2026
Research Briefing is here! https://www.nature.com/articles/s41556-026-01932-w
18 March 2026
Our Nature Cell Biology paper is finally out!! Nature Research Briefing to follow..
https://pubmed.ncbi.nlm.nih.gov/41851271/
01 March 2026
We are delighted to welcome a new PhD student, Abbie Martin, funded by MNDA, to our group!
13 February 2026
Outreach event at High Storrs organised by the Neuroscience Institite. Y11 students loved looking at motor neurons under the microscope, doing fly dissections and learning about fluorescent proteins. Anna, Sabin and Tanya who ran our lab's stall were impressed by the students' knowledge and enthusiasm. Photos in the Gallery.
26 January 2026
Preprint originating from Brittany's thesis work, on STMN2 regulation by stress granules and translation deficits, is out! https://www.biorxiv.org/content/10.64898/2026.01.25.701321v1
6 January 2026
We are happy to welcome a new team member (PDRA) - Katie Gelder!
11-12 December 2025
The team has run a ver successful 1st UK Biomolecular Condensates Network symposium at the Edge, Sheffield, for which TS was a local organiser. We had Tuomas Knowles and Jernej Ule as keynotes and many other excellent researchers speaking. Photos in the Gallery very soon!
 5-7 December 2025
5-7 December 2025TS attended the 36th symposium on ALS/MND in San Diego, where she gave a talk on ALS-FUS and chaired a session.
13 November 2025
Huge congratulations to Brittany for passing her viva with minor corrections! Very grateful to the examiners Zevik Melamed and Alison Twelvetrees.
11 November 2025
TS has been awarded an FLF renewal, with Cara being a named researcher on this project. Happy days!
1 November 2025
The group hosted two really motivated SSC medical students, Ria and Shahid, who worked with Georgia on the DPR optogenetic modeling project - they have now successfully submitted their projects and completed this research attachment course.
31 October 2025
TS attended the 3rd Paraspeckles and condensates conference in Nara, Japan - organised by Tets Hirose and the team. Incredible location, scientific programme and FOOD. See the Gallery.
21 October 2025
The entire team attended ELRIG 2025 Drug Discovery conference in Liverpool. It was very intense two days filled with learning about new technologies and AI developments in drug discovery. Nice dinner at the Botanist Liverpool as well!
12 September 2025
Brittany, Cara and Georgia attended NeuroBioUK - a brilliant annual 1-day conference for ECRs, where Brittany gave a talk and Cara and Georgia presented posters.
9 September 2025
It was very sad to say goodbye to Madeleine - a summer student who wokred with Ved for the last 2 months. It was a pleasure having you in the lab! Nice dinner at the Valencia Crosspool.
5 September 2025
TS presented the lab's work on paraspeckles at the UK condensate network seminar series - check the poster below (Events).
1 September 2025
Preprint on collaborative work by Rahul Samant in Babraham is out! Declining intracellular proteostasis capacity drives misfolded protein secretion in senescent human cells
29 July 2025
SITraN's well represented at the GRC ALS 2025 in Portland ME! TS gave a talk on nuclear TDP-43 condensation and Cara presented a poster on FUS autoregulation (Tobi also gave a short talk).
23 July 2025
Cara's and Ved's paper on novel mechanisms of FUS splicing regulation has been published today in Science Advances: https://www.science.org/doi/10.1126/sciadv.adx1357 Well done both.
22 July 2025
Cara received the best talk prize at the inaugural Nuclei Acids Institute (NAI) symposium held at the Edge, Sheffield. Very well deserved Cara (manuscript will be published tomorrow).
18 July 2025
We are delighted to announce that the 1st UK biomolecular condensate symposium will take place in Sheffield 11-12 December 2025. Please follow the link to the conference webpage.
Register and submit abstract here

3 July 2025
The team is please to welcome a summer placement student Madeleine Fairey - supervised by Ved.
15 June 2025
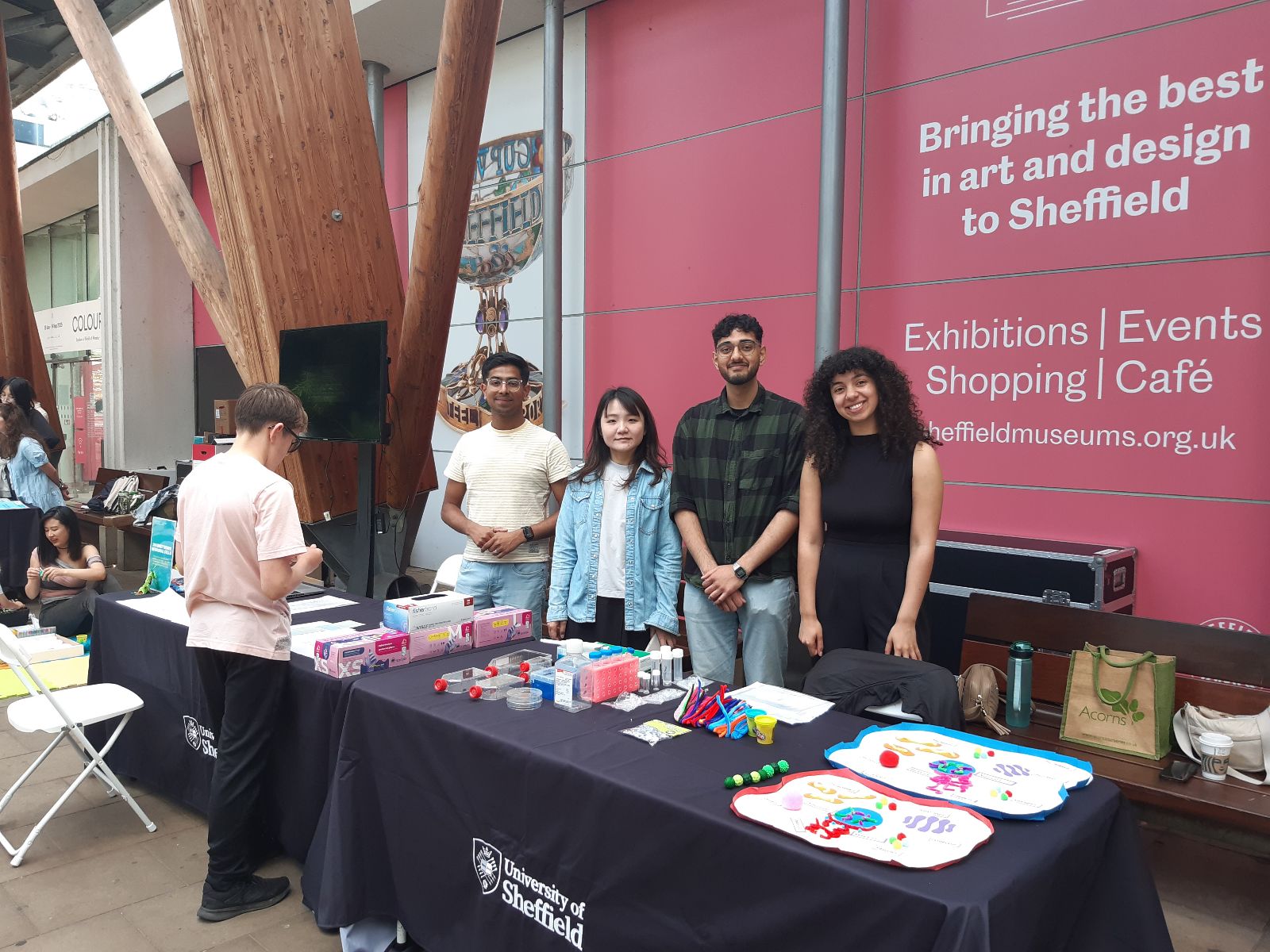
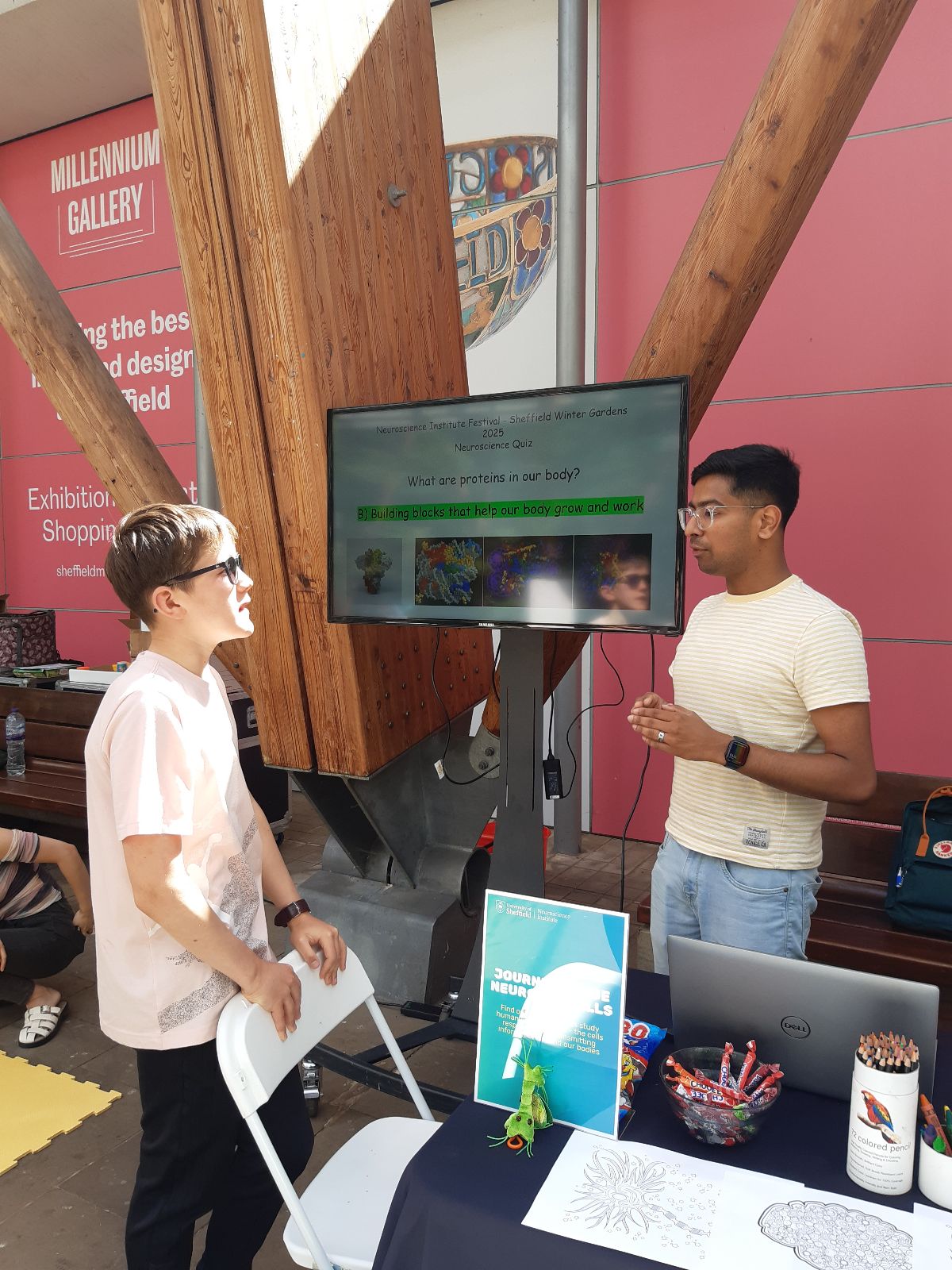
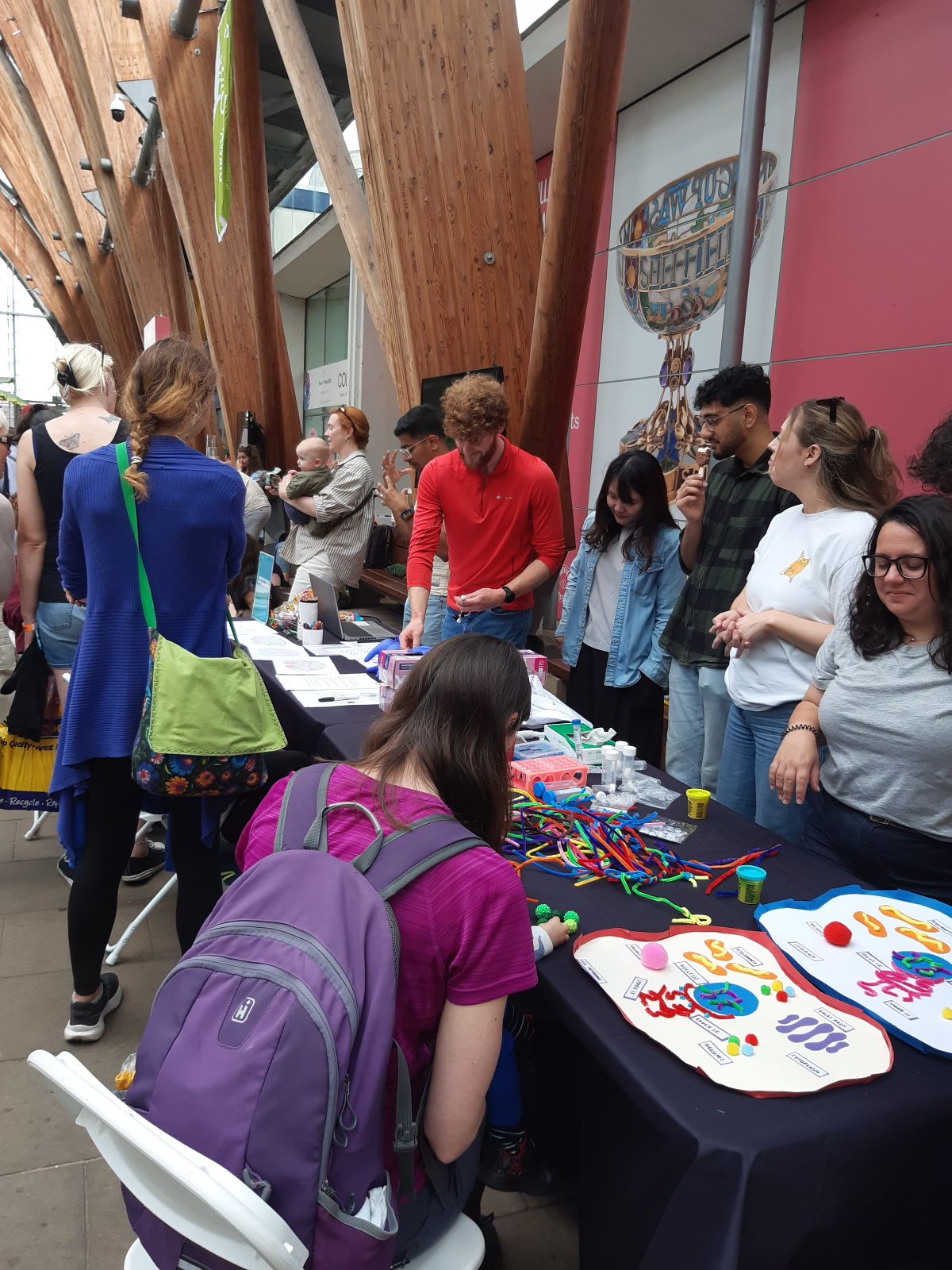
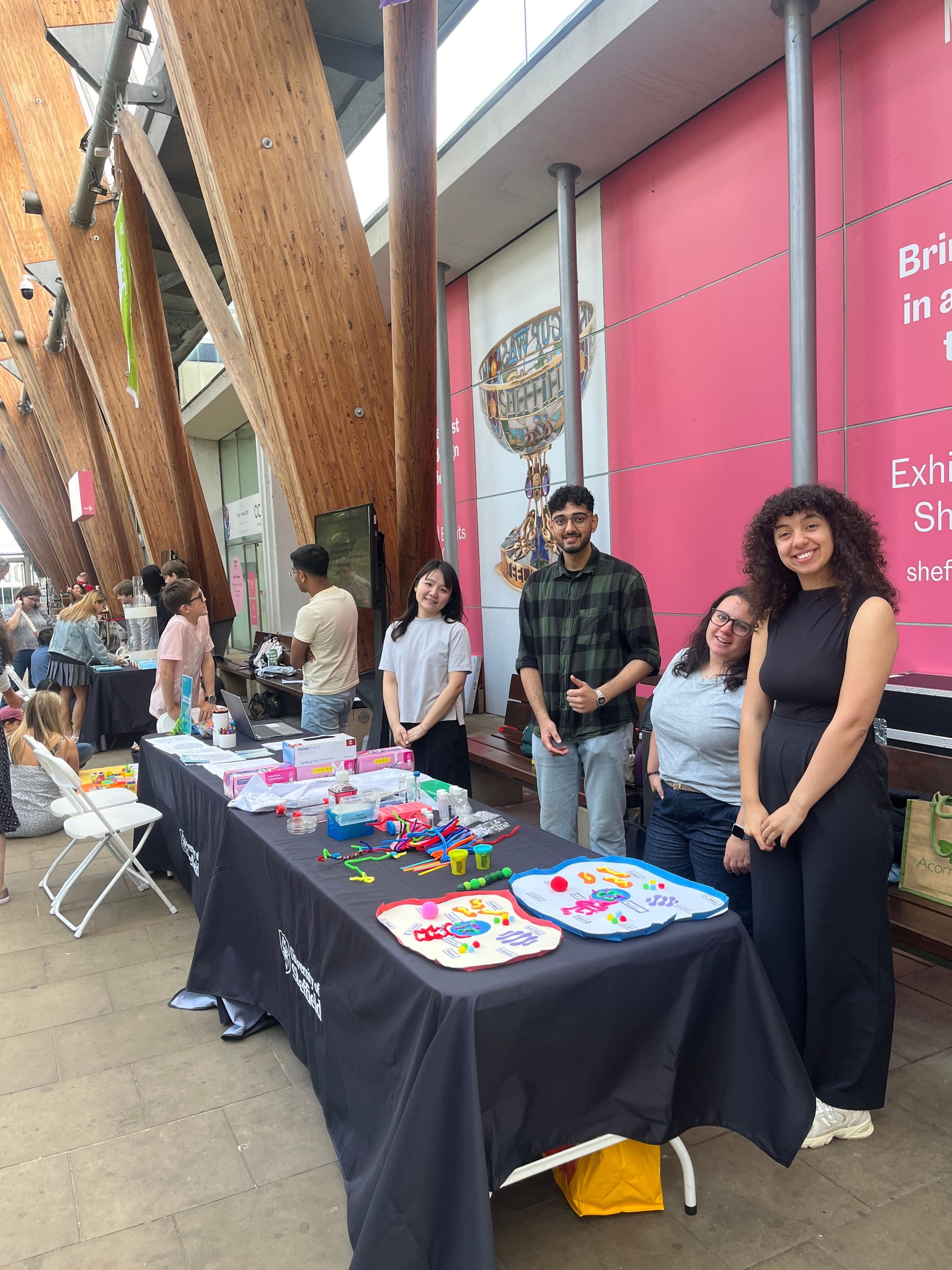
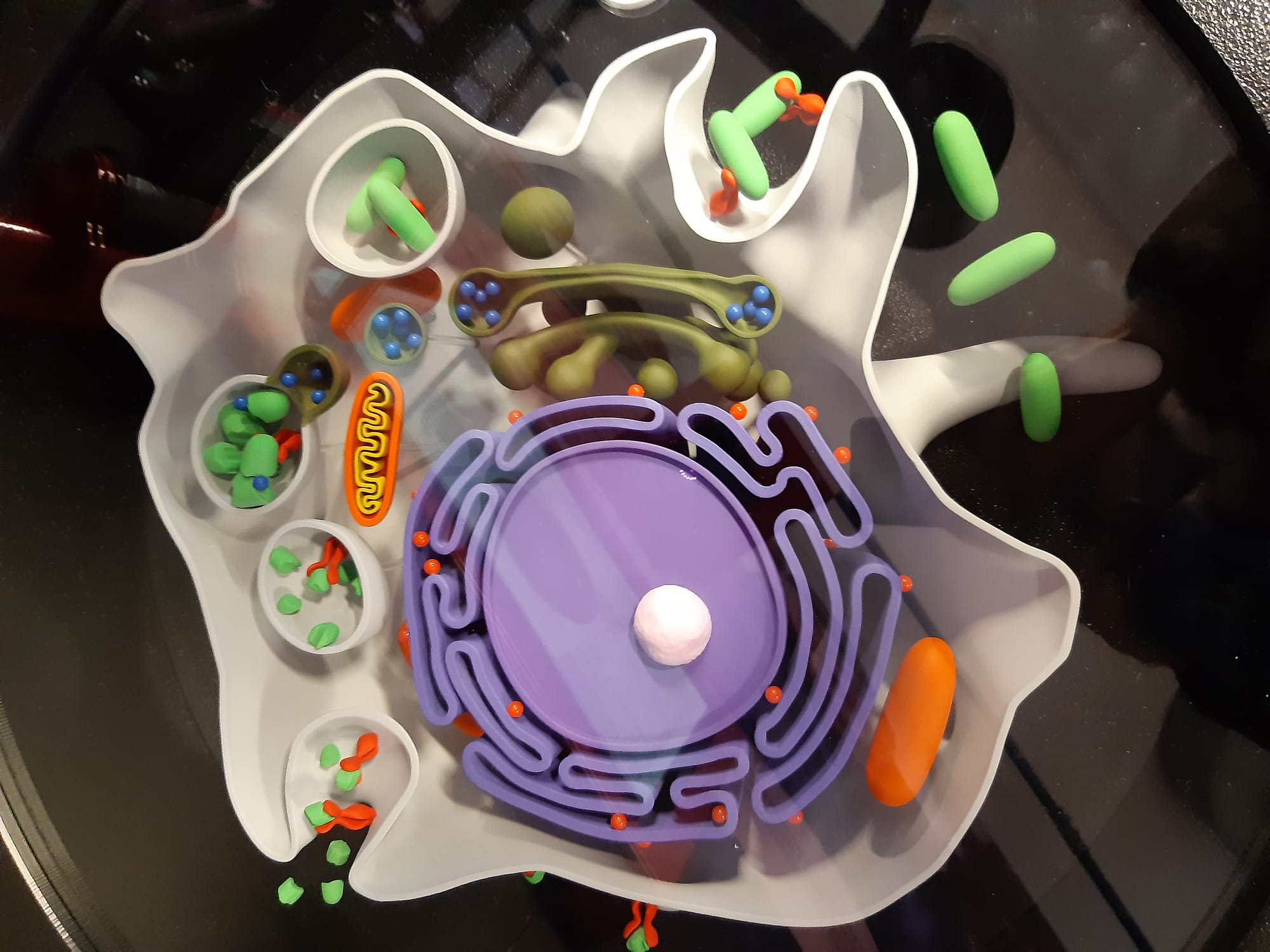
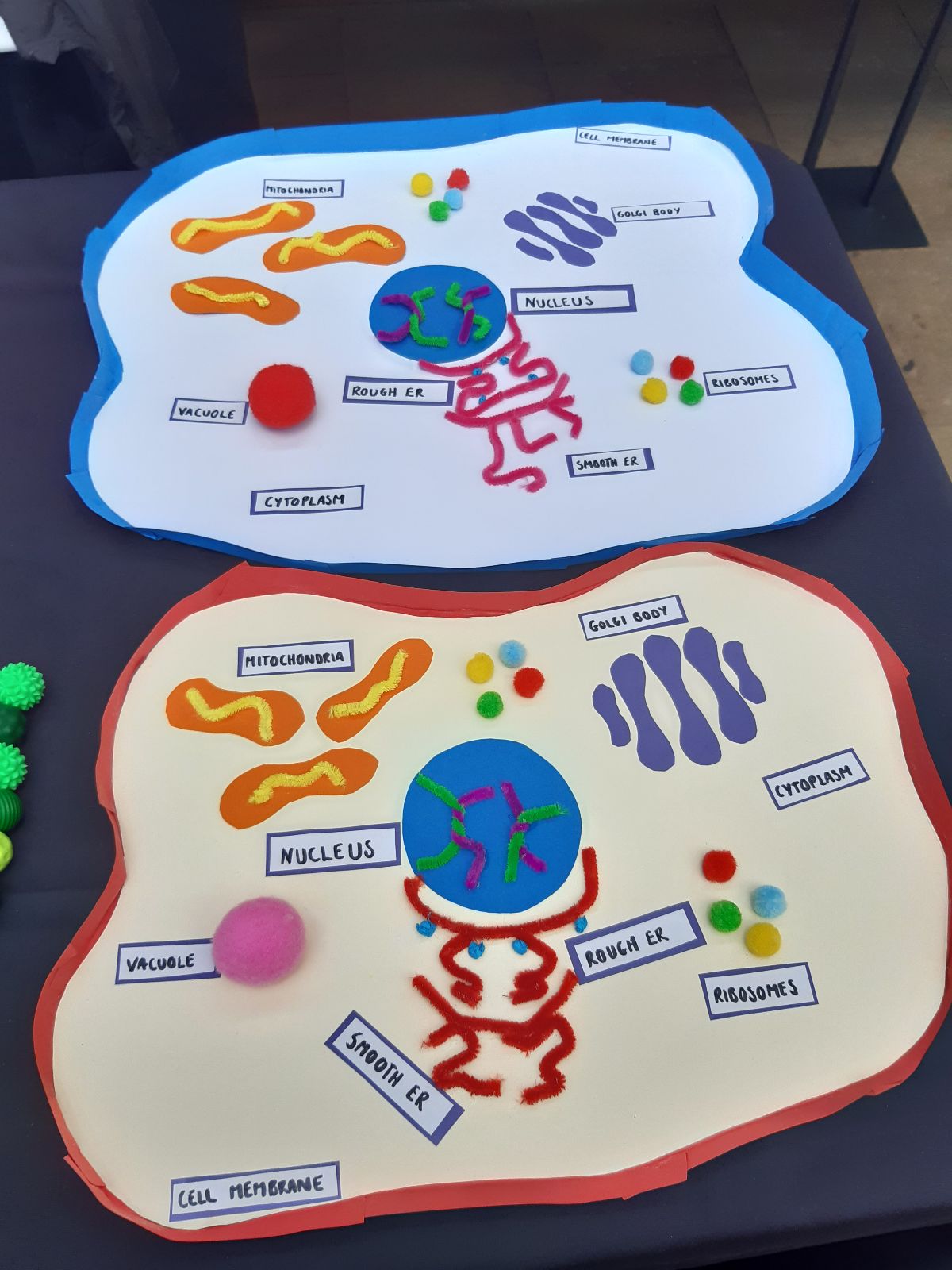
NeuroFest outreach event! Check Gallery-Outreach for photos of Georgia giant cells, molymod and quiz activities. Very popular and fun event - looking to repeat next year!
25 April 2025
Nucleic Acids Institute had its first PI retreat, to celebrate the launch of this cross-cutting collaborative centre: NAI.
17 April 2025
Goodbye dinner at Thyme café Sheffield. Sad goodbye to Emma from Archa's lab and temporary goodbye to Rachel who is off to maternity leave. Emma did everything she planned in the lab (including developing a passion for neurons and organoids), and befriended everyone in the group and beyond. Rachel has submitted her fellowship applications and can now relax (until the baby arrives at least!). Check the Gallery for the hilarious gifts that both received!
Also had Olivier Duss from EMBL over for dinner - who was visiting Sheffield to give a SM@SH talk. Great discussions and busy week!
15 April 2025
We are delighted that we are finalising the agreements on My Mame'5 Doddie Foundation grant - Advancing treatments award and welcoming Fadi Sourakieh on the team. Fadi has stepped in to lead all the chemistry on this project - due to commence on 1st June. Can't wait to start work on small molecule modulators of NEAT1_2/paraspeckles. Check the Foundation portfolio here.
8 April 2025
More conference talks - Sabin and Ved presented at the White Rose RNA Salon 2025 in York - the first external presentations for both. Well done, we've heard you nailed it!
4 April 2025
TS visited Maria Perander's lab in Tromso, Norway - finally (the trip was in the planning for 6 years..). Great seminar and discussions.. and a lot of snow! (check the Gallery).

26 March 2025
Congratulations to Rachel who gave an oral talk (Nuclear TDP-43 pathology in ALS) at the Stem Cell Models of Neurodegeneration 2025 conference (26-28 March, Edinburgh, UK) and also served as a session chair. Emma (visiting PhD student from Archa Fox's lab) and Georgia also presented posters. It was a fantastic conference with great line-up of speakers.

3 February 2025
FUS intron autoregulation manuscript, with Cara and Ved as joint first authors and pretty much everyone in the lab as authors, have been published. Great collaboration with Eugene Makeyev's team at KCL: https://www.biorxiv.org/content/10.1101/2025.02.01.633781v1
25 January 2025
Review in Lancet Neurology on ALS-FUS, co-authored by TS, is online! https://www.sciencedirect.com/science/article/abs/pii/S1474442224005179
10 January 2025
We are happy to welcome Emma Knowling - a visiting student from Archa Fox's lab (UWA) who will study the phase separating behaviour of SFPQ protein in neurons for the next 3 months with us.
12 December 2024
Tanya, Rachel and Georgia attended the London RNA Club Minisymposium on the RNA Biology of the Nervous System, where Tanya gave a talk: https://www.kcl.ac.uk/events/rna-biology-of-the-nervous-system. It was great to see Jacqui - the lab alumnus who is now a postdoc at King's CDN!
14 November 2024
Sheffield Nucleic Acids Institute (NAI) first management board meeting - it is fantastic to be a part of this initiative. Watch for NAI launch day and research symposium updates: https://www.sheffield.ac.uk/nucleic-acids
31 October 2024
Most of the team attended 2nd Paraspeckles (and other condensates) symposium on Rottnest island, Western Australia. Quokkas were of course a highlight ... but the science was excellent too!!
22 October 2024
We are delighted that our project Small molecule hit optimisation for lncRNA NEAT1_2 – a molecular target for neuroprotection in (sporadic) ALS" (collaborative project with Cardiff University) has been supported by My Name’5 Doddie Foundation Advancing Treatments Award. We are excited to continue the development of NEAT1_2 focussed therapeutics.
1st October 2024
We welcome 3 new(ish) PhD students to the lab: Emily Day and Ved Kumar who are not that new (MSs and/or technician in the lab before; Faculty/Neuroscience Institute internal funding) and Georgia Boothe who is joining us from Southampton (MRC DiMeN student). Celebratory sweets are in order at the Friday lab meeting!
25th September 2024
Cara's and Brittany's Cell Reports paper was featured in a Spotlight by Trends in Neurosciences. Congratulations to all the authors on this achievement!!! https://www.cell.com/trends/neurosciences/fulltext/S0166-2236(24)00175-9

25th September 2024
The lab welcomed SSC research attachment medical students who will do a 6-week project on optogenetics, Erynn and Jasmina.
16th September 2024
Rachel presented the data on m6A / FUS crosstalk in ALS-FUS at NeuroBioUK 2024. Excellent conference, co-organised by a SITraN researcher Alison Twelvetrees.
9th September 2024
Sabin has joined the team as a technician on the ARUK funded project working with Rache. Welcome Sabin!!
10th August 2024
Anna, together with other SITraN researchers, run the annual Soak the scientist event in Endcliffe park.
12th July 2024
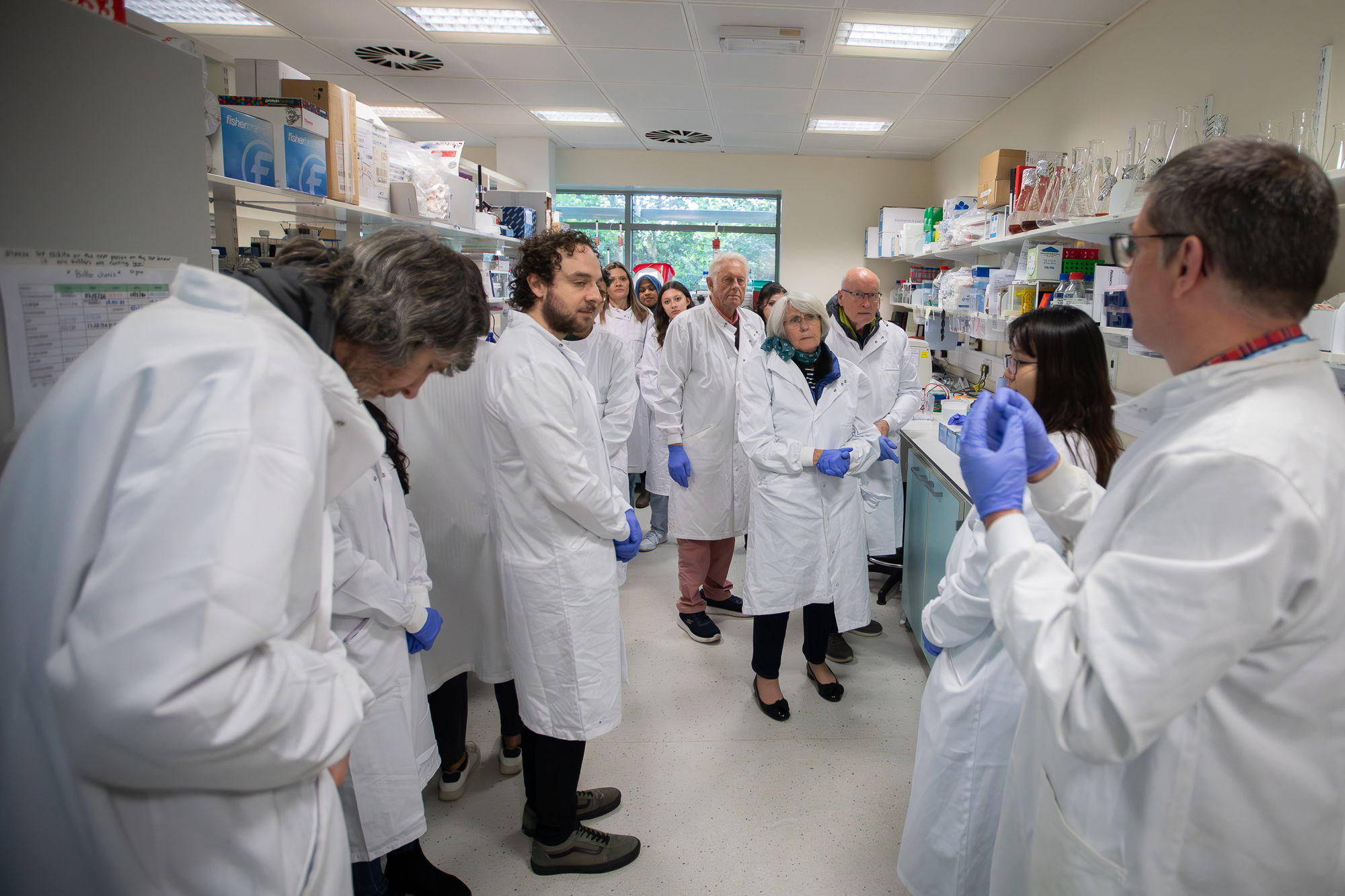
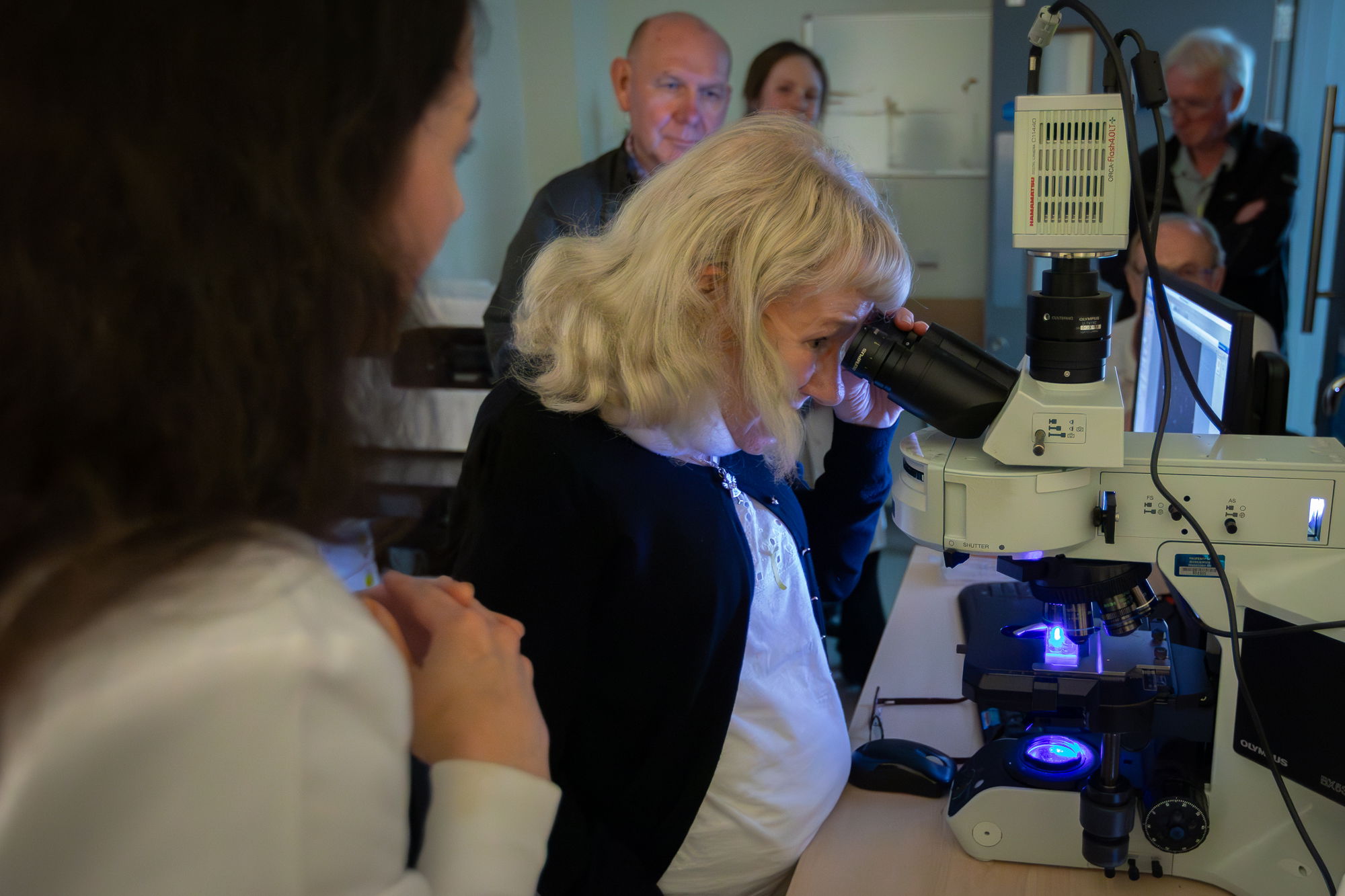
TS lab was involved in organising SITraN open day - which has returned, after 5 years, to attract ~100 members of public for a fun day of activities and talks at the Institute. Rachel and TS ran microscopy session, and Cara manned a molecular biology section slot. Check pics in the Gallery!
3rd July 2024
Rachel gave a talk on translation dysregulation by mutant FUS at Translation UK in Guildford, Surrey.
3rd July 2024
Ruaridh and Rachel's invited review on TDP-43 role in nuclear bodies has been published in Biochem Soc Trans: https://doi.org/10.1042/BST20231447
28th June 2024
Cara and Brittany's Cell reports paper on TDP-43 regulation by nuclear condensation during stress and impact on STMN2 is published! https://doi.org/10.1016/j.celrep.2024.114421
22nd June 2024
Ruaridh presented at Single molecule imaging (SM@SH) seminar on his work using FIDA technology in drug discovery.
15th May 2024
Anna gave an invited talk on nuclear TDP-43 condensation and STMN2 at 11th FTD UK conference in Cambridge.

30 April 2024
We (Archa Fox and myself) are excited to announce that the 2nd Paraspeckle biology conference ("Paraspeckles and other condensates") will take place on Rottnest island (Perth Australia), on 30th October 2024. More information to follow. https://samphirerottnest.com.au/
If you are a paraspeckle/condensate enthusiast and interested to join, please message TS directly.
13 March 2024
Today at the lab meeting three completed manuscripts and a pilot grant awarded to TS and Rachel have been celebrated. The common theme in our work currently is condensates that look like donuts! Hence the donut theme.. Pizza to follow today.. Check the gallery!

 5 March 2024
5 March 2024Lab's second preprint has followed shortly after! https://www.biorxiv.org/content/10.1101/2024.03.05.581933v1 Rachel's and Jess' work on optogenetic modeling of C9-DPR pathology.
20 February 2024
Lab's first preprint posted on SSRN: https://papers.ssrn.com/sol3/papers.cfm?abstract_id=4721338
This work is on the mechanisms of TDP-43 regulated paraspeckle condensation which had been in the making for the last 2 years.
18 February 2024
This time, it was Brittany's turn to present data on nuclear TDP-43 condensation in ALS - collective lab effort, including some of her PhD project data. Manuscript is currently in revision.
29 January 2024
Rachel presented on opto-DPRs at a departmental seminar - this work should be preprinted soon.
25 January 2024
New PhD is advertised in the lab (optogenetics), with deadline on 13th February: https://www.findaphd.com/phds/project/deciphering-the-c9orf72-dpr-aggregation-pathology-in-neurodegeneration-using-optogenetic-and-in-vitro-analyses-of-modifiers/?p168134
10 December 2023
Rachel and TS attended and gave oral talks at 34th Symposium on ALS/MND in Basel (6-8 Dec). It was Rachel's first MND symposium - hope one of many!

9 October 2023
1st paraspeckle symposium was a success!! Photos are in the gallery. Looking to repeat in 2024 in Australia.
1 October 2023
We are delighted to welcome Ruaridh - new PhD student in the lab working on a MNDA-funded project (joint with Scott Allen).
19 September 2023
TS (main supervisor) and Mark Dickman (Engineering, co-supervisor) had their BBSRC DTP PhD project selected. If you are interested to apply for a PhD in TS lab, get in touch - link to project
15 September 2023
Guillaume Haurbergue's paper on PGC1a as an RNA export adapter has been published in Nat Comms. TS is really happy to have contributed to this really nice story! https://pubmed.ncbi.nlm.nih.gov/37679383/
6-8 September 2023
Cara and TS attended the 1st TDP-43 conference in Trieste and presented some of the most recent unpublished data on the role of TDP-43 in paraspeckles (talk and poster). 3 days of TDP-43 biology - what can be better?..
8 September 2023
Brittany has passed her confirmation review with no corrections!
25 August 2023
Emily presented on her MSc thesis. TS was marking the other session so could not attend but everyone said she did fantastic. Congratulations on passing the final bit of your assessment Emily!
19th August 2023
Ved and Anna took part in the Soak a Scientist challenge. Not only got they soaked but Ved also baked for the event. Photos to follow in the gallery.
10th August 2023
Department sponsored trip to Yorkshire Sculpture park! Lots of gaming and sunshine. TS was gutted because she could not go :(
July 2023
It's time to say goodbye to Jess who is off to travelling the world. We wish you all the luck in this new adventure (and please be careful whilst swimming to Cuba - may be take a couple of lessons first!). The image from Jess' mug - also in the gallery! A trip to the Notty house was essential - Jess requested to have one last local pie .. sounded like she'll miss the pies more than us..

17 July 2023
Jess and Cara presenting at the Medical Research School Day 2023:

 July 12-14 2023
July 12-14 2023Ved, Jess, Brittany and Anna attended ENCALS - the first (I am sure of many!) ENCALS for them. Lots of discussions by the posters, fantastic time.

July 2023
TS and Archa Fox (UWA) are organising the first international conference on Paraspeckle biology!!! 8th October 2023, Surrey (satellite to RNA granules). Get in touch if you think your research is relevant: Conference contact/enquiries: paraspeckle.symposium@gmail.com. Rachel and Cara are conference technical committee.
5 July 2023
TS gave an invited talk about her fellowship journey at the Fellowship Day 2023 at the Department of Neuroscience.
01 July 2023
The lab welcomed a new postdoc - Anna Sanchez Avila, who was previously with Chris Henstridge in Dundee and just finished her PhD. Photos to follow!
June 2023
Brittany attended MND EnCouRage UK 2023 and talked about her project to the audience that included patients living with the disease and their carers.
April 2023
Jess, Ved and Brittany had their posters accepted to Encals 2023 in Barcelona so the lab will be well-represented at the conference.
29 March 2023
We are delighted to receive BBSRC Australia partnering award with Archa Fox at UWA. As part of this award, we will organise the 1st international Paraspeckle Biology symposium focused solely on paraspeckles. More detail to follow!
8 March 2023
We have a PhD studentship advertised on findPhD - PTMs in regulation of RNP granules (withMark Dickman and Caroline Evans), international candidates can apply (deadline 5 Apri) - use this link.
27 February 2023
Jess delivered a fantastic presentation at the internal SITraN seminar series. Her talk title was: Using optogenetic tools for modeling neurodegenerative disease pathology. Very well done!
31 January 2023
We bid a sad farewell to Jacqui - who secured a postdoc at Centre for Developmental Neurobiology, King's College London. To celebrate this (not Jacqui's departure but her new exciting appointment), a visit to Nottingham House was paid and a lot of pies were consumed.. Good luck with your new adventure Jacqui! Missing you..

January 2022
We are recruiting! The lab is looking for a postdoctoral researcher with a bioinformatics background, to join the UKRI FLF-funded project. The position is full time and 3 years in duration. If interested, please contact TS directly to discuss by email: t.shelkovnikova@sheffield.ac.uk
20 January 2022
Jacqui and Rachel's work on super-resolution imaging of paraspeckles in cell models of ALS was presented at the SM@SH Single Molecule showcasing event in Sheffield by Rachel.
 15 December 2022
15 December 2022Christmas dinner -Hautbergue and TS labs at the Lescar!

13 December 2022
RNA granules 2023 conference Register interest page is now live! https://www.biochemistry.org/events-and-training/events-calendar/rna-granules-2023/
Together with colleagues from Surrey, Sheffield and Manchester and support from the Biochemical Society, we are organising the international conference “RNA Granules 2023: from basic biology, functions to drug discovery", to be held in Surrey (UK), 9-11th October 2023.
 The previous RNA granules conferences held in 2014 and 2017 in Canada and Germany were a success (the 2020 Vancouver meeting was cancelled due to COVID). The 2023 conference sessions will cover several themes, including Biocondensates Biogenesis and Dynamics, Biocondensates in Neurological Disorders, Cancer and Infections, Innovation and Methodological Advances, Therapeutics, Physico-Chemical properties of Phase Separation. Roy Parker is a confirmed keynote speaker. More information is to follow in early 2023 - stay tuned by signing up to the conference updates using the link above!
The previous RNA granules conferences held in 2014 and 2017 in Canada and Germany were a success (the 2020 Vancouver meeting was cancelled due to COVID). The 2023 conference sessions will cover several themes, including Biocondensates Biogenesis and Dynamics, Biocondensates in Neurological Disorders, Cancer and Infections, Innovation and Methodological Advances, Therapeutics, Physico-Chemical properties of Phase Separation. Roy Parker is a confirmed keynote speaker. More information is to follow in early 2023 - stay tuned by signing up to the conference updates using the link above!12 November 2022
TS's profile on ResearchGate achieved 10,000 reads!
November 2022
The lab welcomed 2 new postdocs, Cara and Rachel - welcome to the team!
October 2022
TS's first MSc students at Sheffield, Ved and Jess - who are have joined as technicians now - had their graduation ceremony! So pleased and proud for them!!

October 2022
The lab welcomed a PhD student, Brittany, and three technicians, Ved, Jacqui and Jess - warm welcome!
29 September 2022
Our review on the use of OTIs as precision gene editors in neurodegenerative diseases led by Ben Bax (Cardiff) has been published: https://www.mdpi.com/1422-0067/23/19/11541
13 September 2022
Our collaborative MRC/AZ CLD project (with Prof John Atack at Cardiff MDI) was awarded! We will be granted access to the AZ small molecule collection to find NEAT1/paraspeckle targeting molecules.
26 August 2022
Our Nucleic Acids Research paper on therapeutic targeting of NEAT1/paraspeckles has been accepted!! See here (and soon in the paper section).
5 April 2022
TS and Guillaume Hautbergue were awarded a 3-year MRC research grant to establish a biochemical assay for analysis of ALS-linked RNP complexes, suitable for screening of small molecules. The recruitment on this grant (3-year postdoctoral position, full time) to begin soon (will be posted here).
4 February 2022
TS and Kurt De Vos have been awarded a PhD studentship to identify drugs that modulate STMN2 splicing, by MND Scotland. Needless to say that we are honored and excited to be selected within this completive scheme. The project is currently advertised on Findaphd, until 6th May:
https://www.findaphd.com/phds/project/small-molecule-modulators-of-alternative-splicing-as-a-novel-therapy-for-amyotrophic-lateral-sclerosis-als/?p143014
6 September 2021
Lab has moved to SITraN, Sheffield. Exciting times!!
5 May 2021
Tatyana has been awarded a BBSRC grant, together with Nicolas Locker from Surrey, to look into the molecular mechanisms of stress granule - paraspeckle crosstalk.
16 March 2021
MND Research blog about our work was published, check it here: https://mndresearch.blog/2021/03/16/flexible-molecules-and-droplets-researching-and-targeting-rna-protein-complexes-in-mnd/
02 March 2021
Our study on the analysis of FUS aggregate composition is now available as a preprint: https://www.biorxiv.org/content/10.1101/2021.03.02.433611v1
11 January 2021
The paper with our collaborators in Tokyo, Nevada and Moscow has been accepted for publication at the RNA Biology journal.
08 December 2020
Tatyana is this year's winner of the ENCALS Young Investigator Award!
07 October 2020
The lab has received a grant from the Welsh Government for a COVID-19 drug discovery project. Can't wait to start it!
04 May 2020
Our paper on neurological phenotypes in Neat1 knockout mice (previously available as preprint), with Haiyan, Camille and Tatyana as authors, has been accepted for publication in the Translational Psychiatry journal.
30 April 2020
Our paper on the effect of FUS frameshift mutations, with Haiyan, Camille and Tatyana as authors, has been accepted for publication in the Molecular Brain journal.
12 February 2020
Tatyana presented at CU School of Dentistry seminar: “Therapeutic targeting of RNA – expanding the boundaries of the druggable genome?”
14 January 2020
Tatyana invited to give a talk on paraspeckle regulation by stress granules at the RNA UK 2020 meeting
https://www.rnauk2020.org/
21 November 2019
Invited commentary on our J Cell Biol paper was published in Cell Stress
check free text: https://www.cell-stress.com/researcharticles/2019a-an-microbial-cell/
23 October 2019
Haiyan had her PhD viva – only minor corrections! Congratulations!!!
23 September 2019
The very first lab’s preprint, with Haiyan, Camille and Tatyana as authors, published on bioRxiv: https://www.biorxiv.org/content/10.1101/773333v1
Contact information
Stay in touch
Contact
Internal Use
-80C Chemical Data Base
Includes description and location of -80C contents
Events
3rd Paraspeckles and condensates conference
Info here: https://www.ukcondensates.co.uk/monthly-network-seminar-series
Paraspeckles and other condensates conference
1st Paraspeckle Biology conference